
Specialization:
Education:
Dr. Reich received his Ph.D. from the University of California, San Francisco in 1984. After an NIH post-doctoral fellowship at UCSF, he joined the faculty at UCSB in 1987. His awards include: a Regent's Junior Faculty Fellowship (1987); an American Cancer Society Faculty Research Award (1991); and the UC President's Award for Excellence in Undergraduate Research (1994).
Research:
Reich Lab Website
Research Objectives:
We are interested in the study of enzyme function, primarily at the levels of catalytic and specificity mechanisms, with a long term goal of developing enzyme inhibitors with potential therapeutic application. The current focus is on both bacterial and mammalian enzymes involved in the epigenetic process of DNA methylation and on telomerase. We also work on the development of new methods and technologies to investigate cell-biological, biochemical and biomedical questions.
![Text Box: Figure 1: [A] Bacterial DNA cytosine methyltransferase M.HhaI complexed to DNA (yellow) with the catalytic loop in the open (red) and closed (blue) conformations. The conformational transition has been traced using fluorescence and NMR methods, and forms a key part of an induced fit mechanism whereby sequence recognition drives base flipping and loop movement. [B] and [C] show highlights of the recognition interactions.](http://chem.ucsb.edu/sites/secure.lsit.ucsb.edu.chem.d7-3/files/sitefiles/people/reichn/index_0clip_image002.gif)
DNA methyltransferases: epigenetics, enzyme mechanisms, and the development of novel therapeutics
S-Adenosyl methionine-dependent DNA methyltransferases modify DNA sequence-specifically. Bacterial DNA is modified at both cytosine (N4 or C5) and adenine (N6), while mammalian DNA is modified only at cytosine C5. The bacterial enzymes are involved in restriction/modification, mismatch repair, replication timing, and gene regulation. These enzymes are essential to the virulence of several human pathogens. Numerous high resolution x-ray structures (see Figures 1, 2) are available for these bacterial enzymes, enabling our efforts to understand the following aspects:
- Mechanisms of catalysis
- Importance of conformational mechanisms, including DNA bending, base flipping, intercalation, and protein loop motion towards catalysis and specificity.
- Antibiotic design based on inhibiting the bacterial DNA MTases.
- The engineering of new specificities through the combined use of unnatural amino acids and in vitro evolution.
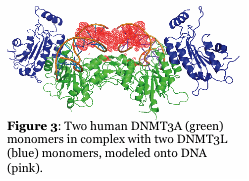
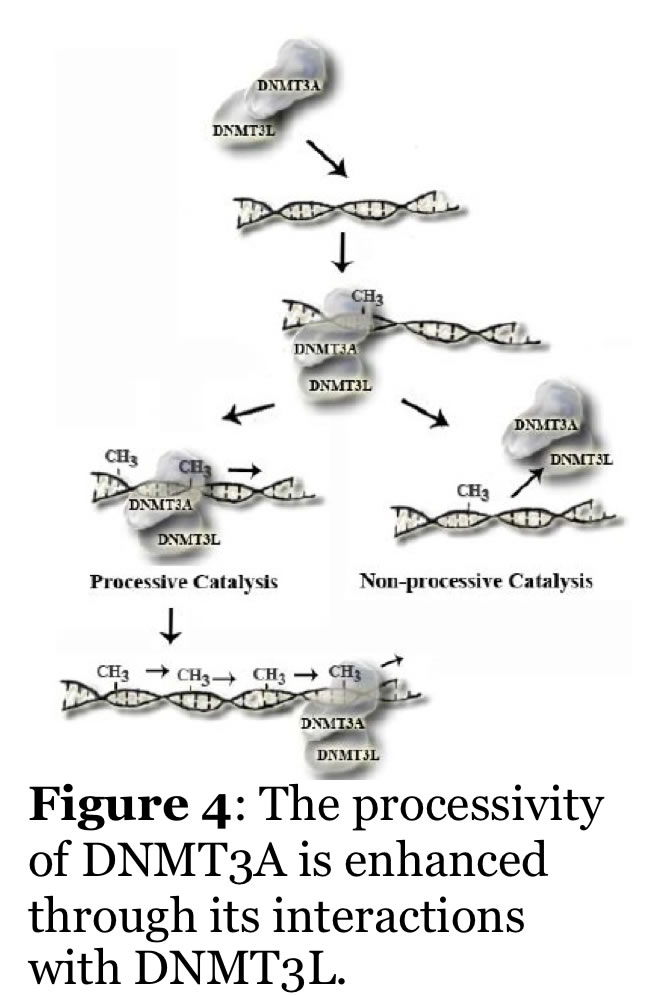
 Mammalian DNA methylation is involved in parent of origin imprinting, x-chromosome inactivation, host defense, and tissue-specific gene regulation. Aberrant DNA methylation of tumor suppressor genes is a primary and early event in human tumorigenesis. Mutations in the DNA cytosine methyltransferases (e.g., DNMT3A, Figure 2) are detected in patients with various forms of leukemia. Similarly, proteins that regulate DNMT function, such as the helicase Lsh, and the transcription factor cMyc, alter DNMT function. A major unresolved question is how do the patterns of DNA methylation, which occur in the CpG dinucleotide, become established? Recent work shows that the mammalian DNA cytosine methyltransferases (Dnmt1, Dnmt2, Dnmt3 family) interact with other cellular proteins, and that de novo DNA methylation is directed in part by non-coding RNA. Our efforts are focused on studying the human enzymes to understand how interactions with other proteins and RNA direct the function and specificity of these enzymes. For example, we recently showed that the non-catalytic DNMT3L activates DNMT3A function by enhancing its ability to methylate DNA with multiple CG sites (processive catalysis, Figure 4). This work includes the human and mouse enzymes, human cancer cells, and embryonic stem cells. We have also identified inhibitors of the mammalian enzymes which are cell active.
Mammalian DNA methylation is involved in parent of origin imprinting, x-chromosome inactivation, host defense, and tissue-specific gene regulation. Aberrant DNA methylation of tumor suppressor genes is a primary and early event in human tumorigenesis. Mutations in the DNA cytosine methyltransferases (e.g., DNMT3A, Figure 2) are detected in patients with various forms of leukemia. Similarly, proteins that regulate DNMT function, such as the helicase Lsh, and the transcription factor cMyc, alter DNMT function. A major unresolved question is how do the patterns of DNA methylation, which occur in the CpG dinucleotide, become established? Recent work shows that the mammalian DNA cytosine methyltransferases (Dnmt1, Dnmt2, Dnmt3 family) interact with other cellular proteins, and that de novo DNA methylation is directed in part by non-coding RNA. Our efforts are focused on studying the human enzymes to understand how interactions with other proteins and RNA direct the function and specificity of these enzymes. For example, we recently showed that the non-catalytic DNMT3L activates DNMT3A function by enhancing its ability to methylate DNA with multiple CG sites (processive catalysis, Figure 4). This work includes the human and mouse enzymes, human cancer cells, and embryonic stem cells. We have also identified inhibitors of the mammalian enzymes which are cell active.
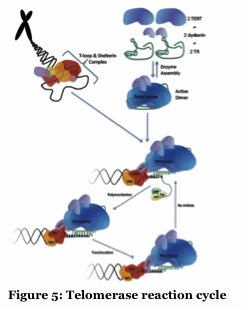
Telomerase extends the ends of chromosomes and displays unique mechanisms of DNA polymerization (Figure 5). We are collaborating with Tom Pettus (UCSB) to exploit known and designed small molecule telomerase inhibitors to better understand the human and Tetrahymena enzymes. Our long term goal is to identify potent and selective inhibitors.
 New tools for cell biology, proteomics, and drug delivery
New tools for cell biology, proteomics, and drug delivery
We are developing three platform technologies with broad basic research and biomedical applications.
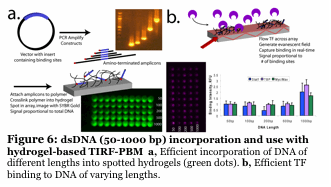
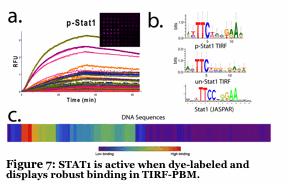
We have developed a unique fluorescence-based method to simultaneously follow the binding of 5-10 individual proteins to a microarray made up of DNA promoters. The DNA is imbedded into a hydrogel, placed onto a slide and the proteins (dye labeled transcription factors) flow across the surface in a microfluidic device. By using unique fluorescent dyes, we observe binding without any wash steps (Figure 6). We determine binding and kinetic constants for thousands of sequences simulutaneously, all while tracking individual proteins. No current technique can do this. We are using this to understand how 3-10 transcription factors work together to form a multiprotein complex of high specificity (Figure 7). Also, we are using this to identify small molecule inhibitors of human transcription factors (e.g., STAT1,3) which are involved in cancer.
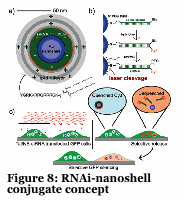
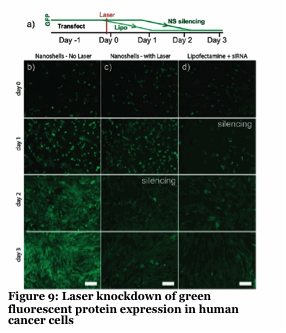
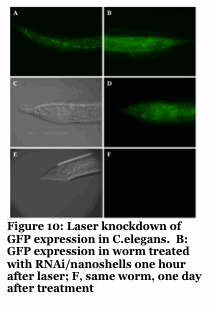
We have developed a gold nanoshell technology which relies on near infra-red lasers to release RNAi, drugs and proteins within human cells and model organisms (Figure 8-10). Nanoshells provide a route to protect the cargo (e.g., RNAi) from degradation. Furthermore, by decorating the particles with targeting peptides we can direct the localization of the material into cells and into organisms. We are collaborating with Dr. Erkki Ruoslahti to target the nanoshells to tumors and atherosclerotic plaques to alter the pharmacokinetics of the drug (e.g., RNAi).
Our approach provides unprecedented spatio-temporal control with a non-harmful laser. The use of this to knockdown specific genes in organisms like C.elegans with single cell and developmentally staged resolution is unprecedented and an ongoing effort.
Publications:
Selected Research Publications
Pollak, A.J., Reich, N.O. 2012. Proximal recognition sites facilitate intrasite hopping by DNA adenine methyltransferase: mechanistic exploration of epigenetic gene regulation. The Journal of biological chemistry, 287(27), 22873-81.
Matje, D.M., Reich, N.O. 2012. Molecular drivers of base flipping during sequence-specific DNA methylation. Chembiochem : a European journal of chemical biology, 13(11), 1574-7.
Laurence, T.A., Braun, G.B., Reich, N.O., Moskovits, M. 2012. Robust SERS enhancement factor statistics using rotational correlation spectroscopy. Nano letters, 12(6), 2912-7.
Bonham, A.J., Wenta, N., Osslund, L.M., Prussin, A.J., 2nd, Vinkemeier, U., Reich, N.O. 2012. STAT1:DNA sequence-dependent binding modulation by phosphorylation, protein:protein interactions and small-molecule inhibition. Nucleic acids research.
Cohn, E.P., Wu, K.L., Pettus, T.R., Reich, N.O. 2012. A new strategy for detection and development of tractable telomerase inhibitors. Journal of medicinal chemistry, 55(8), 3678-86.
Holz-Schietinger, C., Matje, D.M., Reich, N.O. 2012. Mutations in DNA methyltransferase (DNMT3A) observed in acute myeloid leukemia patients disrupt processive methylation. The Journal of biological chemistry, 287(37), 30941-51.
Holz-Schietinger, C., Reich, N.O. 2012. RNA modulation of the human DNA methyltransferase 3A. Nucleic acids research, 40(17), 8550-7.
Holz-Schietinger, C., Matje, D.M., Harrison, M.F., Reich, N.O. 2011. Oligomerization of DNMT3A controls the mechanism of de novo DNA methylation. The Journal of biological chemistry, 286(48), 41479-88.
Matje, D.M., Coughlin, D.F., Connolly, B.A., Dahlquist, F.W., Reich, N.O. 2011. Determinants of precatalytic conformational transitions in the DNA cytosine methyltransferase M.HhaI. Biochemistry, 50(9), 1465-73.
Menezes, S., Gaston, K.W., Krivos, K.L., Apolinario, E.E., Reich, N.O., Sowers, K.R., Limbach, P.A., Perona, J.J. 2011. Formation of m2G6 in Methanocaldococcus jannaschii tRNA catalyzed by the novel methyltransferase Trm14. Nucleic acids research, 39(17), 7641-55.
Vallee-Belisle, A., Bonham, A.J., Reich, N.O., Ricci, F., Plaxco, K.W. 2011. Transcription factor beacons for the quantitative detection of DNA binding activity. Journal of the American Chemical Society, 133(35), 13836-9.
Determinants of Precatalytic Conformational Transitions in the DNA Cytosine Methyltransferase M.HhaI, Matje DM, Coughlin DF, Connolly BA, Dahlquist FW, Reich NO.
Biochemistry. 2011 Jan 13. [Epub ahead of print]
The inherent processivity of the human de novo methyltransferase 3A (DNMT3A) is enhanced by DNMT3L, Holz-Schietinger C, Reich NO. J Biol Chem. 2010 Sep 17;285(38):29091-100.
Escherichia coli DNA adenine methyltransferase: intrasite processivity and substrate-induced dimerization and activation, Coffin SR, Reich NO., Biochemistry. 2009 Aug 11;48(31):7399-410.
Laser-Activated Gene Silencing via Gold Nanoshell-siRNA Conjugates, Braun GB, Pallaoro A, Wu G, Missirlis D, Zasadzinski JA, Tirrell M, Reich NO., ACS Nano. 2009 Jun 15. [Epub ahead of print].
Coupling sequence-specific recognition to DNA modification, Estabrook RA, Nguyen TT, Fera N, Reich NO., J Biol Chem. 2009 Aug 21;284(34):22690-6. Epub 2009 Jun 4.
Tracking transcription factor complexes on DNA using total internal reflectance fluorescence protein binding microarrays, Bonham AJ, Neumann T, Tirrell M, Reich NO., Nucleic Acids Res. 2009 Jul;37(13):e94. Epub 2009 May 31.